Spatial mapping and contextualization of axon subtypes innervating the long bones of C3H and B6 mice
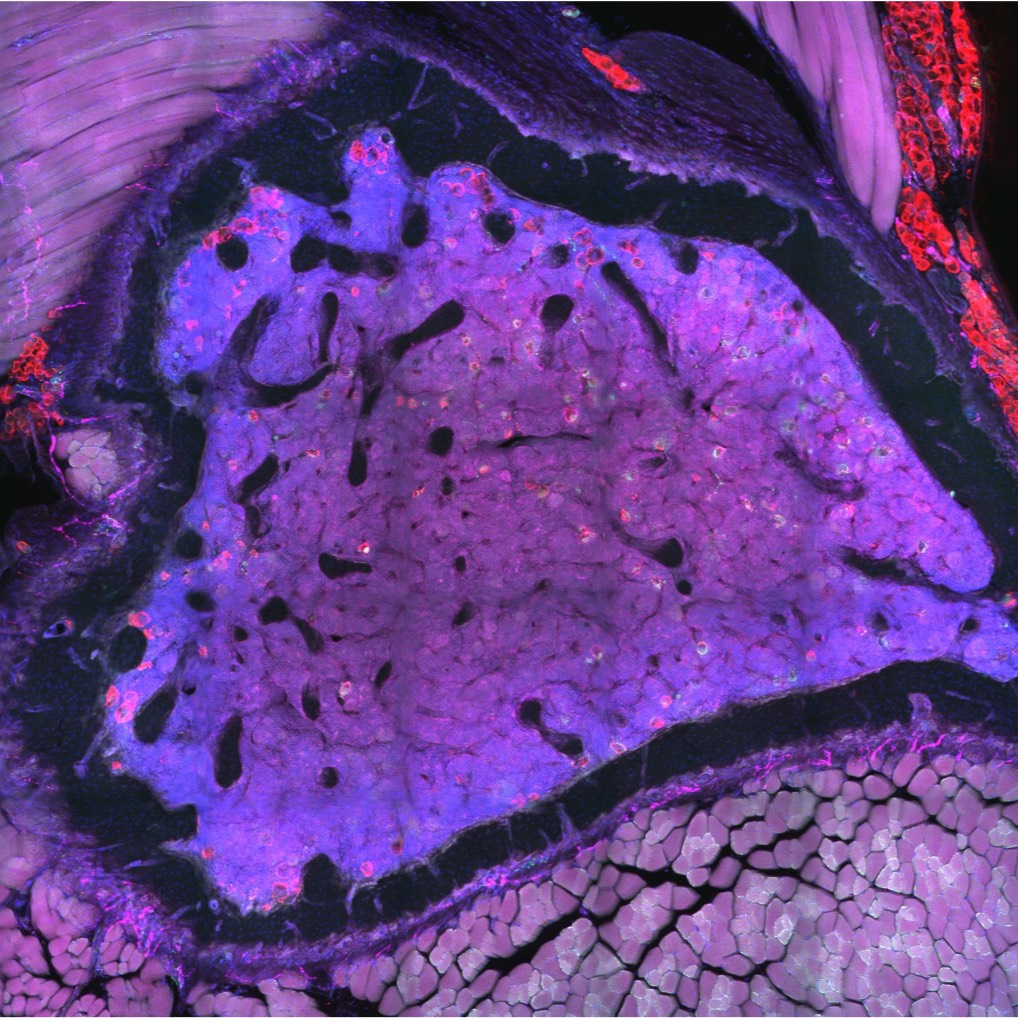
A dataset containing high-resolution confocal images of tranverse cross-sections of the C57BL/6J and C3H/HeJ femur and tibia that have been stained with anti-CGRP (sensory nerve), anti-TH (sympathetic nerve), anti-perilipin (adipocytes), and dapi (nuclei)
Dataset Overview
Study Purpose: Nerves in bone play well-established roles in pain and vasoregulation and have been associated with the progression of skeletal disorders including osteoporosis, fracture, arthritis, and tumor metastasis. However, isolation of the region-specific mechanisms underlying these relationships is limited by our lack of precise maps of skeletal innervation. We developed an optimized workflow for imaging of nerve axons in and around the bone, resulting in novel, comprehensive maps of sympathetic adrenergic and sensory peptidergic axons within and around the full length of the femur and tibia in two strains of mice (B6 and C3H).
Data Collection: This dataset contains confocal images of transverse cross-sections of the C57BL/6J and C3H/HeJ femur and tibia that have been stained with anti-CGRP (sensory nerve), anti-TH (sympathetic nerve), anti-perilipin (adipocytes), and dapi (nuclei). Serial tiled images were taken with a 10x objective on a Nikon spinning disk confocal microscope (µm/px = 0.650, step size = 2.5 µm, number of steps = 21). Stitching of the nd2 multipoint files was done using the FIJI Stitching plugin “snake by rows”. Images are stored both as full z-stack TIFF files and maximum-intensity projection TIFF files and are viewable in FIJI or any other compatible image processing/viewing software.
Primary Conclusion: The innervation of the skeleton is not uniform. Our neuroskeletal maps demonstrate that peripheral axon subtype-specific densities and morphology vary by bone compartment and site and are influenced by mouse genetic strain. While the endosteum is entirely aneural, three distinct patterns of periosteal innervation are evident at sites that are important for bone health, including aneural, anchored/diving, and fascial/parallel patterned surfaces. Ultimately, this work is intended to serve as a guide during the design, implementation, and interpretation of future neuroskeletal studies.
Curator's Notes
Experimental Design: Mice were anesthetized with Ketamine/Xylazine and perfused through the left ventricle of the heart with 10 mL phosphate buffered saline followed by 10 mL 10% neutral buffered formalin (NBF, Fisher Scientific 23-245684). For all experiments, collected tissues were post-fixed in 10% NBF for 24 hours. After fixation, tissues were washed for 2 hours in diH2O. Bones were fully decalcified in 14% EDTA (Sigma-Aldrich E5134), pH 7.4 and equilibrated in 30% sucrose solution prior to embedding in OCT mounting media (Fisher HealthCare 23-730-571). Embedded tissues were cut at 50 µm on a cryostat (Leica). For immunostaining, cut sections on Colorfrost Plus glass slides (Fisher Scientific 12-550-18) were blocked in 10% donkey serum in TNT buffer prior to incubation for 48 hours with primary antibodies at 4°C. After washing 3x 5 min, secondary antibodies in TNT buffer were applied for 24 h at 4°C. The sections were then washed 3x 5 min in TNT buffer, incubated in DAPI (Sigma-Aldrich) for 5 min, and washed again prior to mounting with Fluoromount-G (Thermo Fisher Scientific 00-4958-02). Serial tiled images were taken with a 10x objective on a Nikon spinning disk confocal microscope (µm/px = 0.650, step size = 2.5 µm Number of steps = 21).
Completeness: Complete
Subjects & Samples: Representative transverse sections at 11 longitudinally-spaced levels in the tibia and 6 levels in the femur were chosen for cohorts of C57BL/6J and C3H/HeJ mice at 12 weeks of age for a total of 34 imaging datasets. These samples represent 22 sections of the mouse tibia (11 C57BL/6J, 11 C3H/HeJ) and 12 sections of the femur (6 C57BL/6J, 6 C3H/HeJ).
Primary vs derivative data: The primary folder contains microscopic images (TIFF) from each sample in the study. The derivative folder contains maximum-intensity projections of each full z-stack (TIFF). Max projections were created in and can be viewed in FIJI.
Files
1 - 0 of 0 files
About this dataset
Publishing history
Cite this dataset
Tags
References
Is Supplemented by
Scheller, E., Lab, S., M Brazill, J., & Lorenz, M. (2020). Spatial mapping and contextualization of axon subtypes innervating the long bones of C3H and B6 mice v1. https://doi.org/10.17504/protocols.io.bqu2mwye
Described by
Lorenz, M. R., Brazill, J. M., Beeve, A., Shen, I., & Scheller, E. L. (2020). A neuroskeletal atlas of the mouse limb. https://doi.org/10.1101/2020.09.18.303958
