Pig whole-body with embedded organs using automatic workflow for inserting the organs
This dataset includes lung and heart inserted into the pig whole-body using an automated workflow.
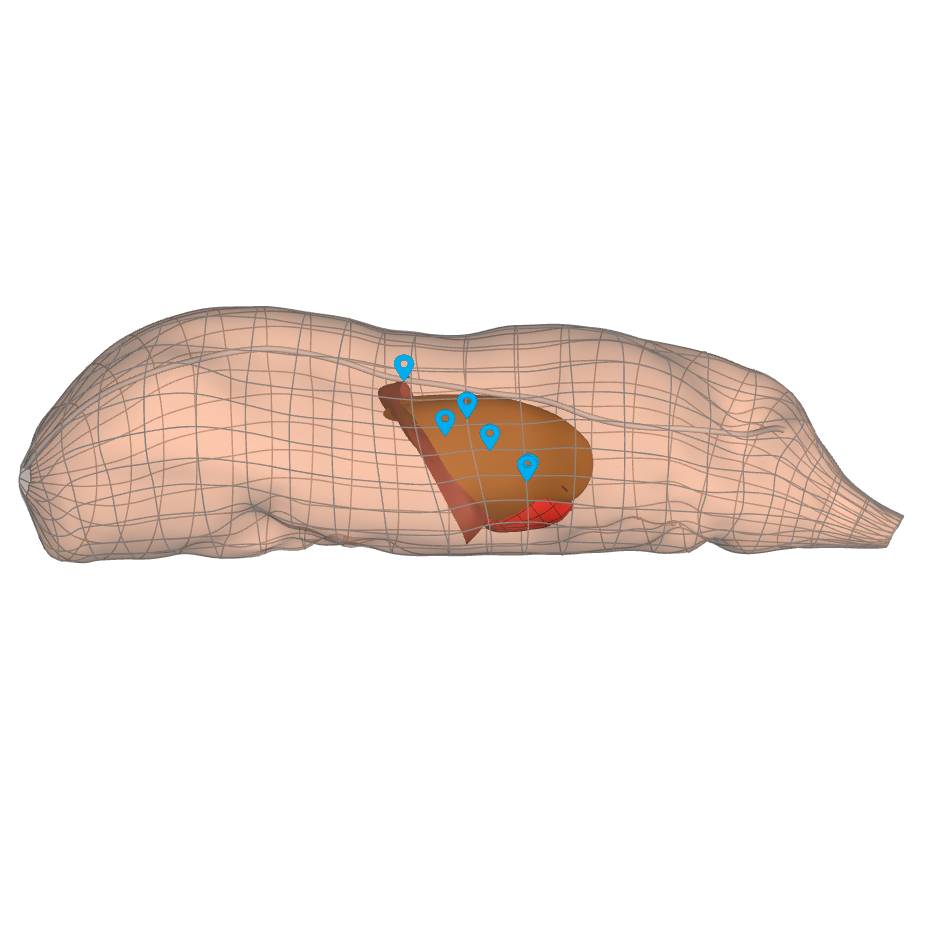
Dataset Overview
Study Purpose: The goal of this work is to automate the pig insertion of 3D organ scaffolds into the whole-body scaffolds and produce pig whole-body with embedded organs.
Data Collection: A MAP Client workflow was created with a new step (organinserter) in python language to automate the insertion of 3D organs into the whole-body scaffolds. The whole-body scaffold along with the organs are passed to the workflow and then it finds the common fiducial markers in the whole-body and each organ. These fiducials then are used to register the organ into the body. The embedded organs include lungs and heart.
Primary Conclusion: None stated
Curator’s Notes
Experimental Design: Not applicable.
Completeness: The study is ongoing and potentially will link to other datasets where the data is used for mapping onto the scaffold.
Subjects & Samples: The automated workflow is not subject/sample-specific and the resultant output is produced for pig models.
Primary vs derivative data: In the primary folder, nodal information and element connectivity of the scaffold are formatted as an exf file. Using the commands in the cmgui file, the scaffold can be visualized using open source software CMGUI. The derivative folder contains JSON files that are used to generate a visualization of the scaffold on the web portal.
Code Availability: https://github.com/ABI-Software/scaffoldmaker (ABI-Software/scaffoldmaker Anatomical scaffold generator using OpenCMISS), https://models.physiomeproject.org/workspace/82f (scaffold source files and workflow to reproduce the files)
Files
1 - 0 of 0 files
