Generic pig stomach scaffold
Annotated pig stomach scaffold available for registration of segmented neural anatomical-functional mapping of enteric neural circuits.
Dataset Overview
Study Purpose: The goal of this work is to create an annotated generic pig stomach scaffold for the registration of segmented data obtained by experimental groups.
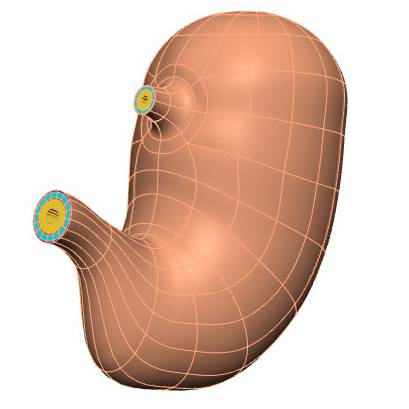
Data Collection: An anatomically-based 3D scaffold of the pig stomach is created to map nerve ending pathways. The stomach scaffold is generated with a configurable central path defined from the fundus apex to the duodenum. A user can annotate the fundus, body, antrum, pylorus, and duodenum along the central path and use that to define the respective regions of the stomach. The cross axes of the central path provide control of the major and minor radii of the stomach in different sections. The pig stomach scaffold is parameterized with data to represent the anatomy as accurate as possible. Four wall layers (mucosa, submucosa, circular muscle, and longitudinal muscle) are added to the pig stomach scaffold. The outer surface of the scaffold is annotated as the serosa.
Primary Conclusion: None stated
Curator’s Notes
Experimental Design: Not applicable.
Completeness: The study is ongoing and potentially will link to other datasets where the data is used for mapping onto the scaffold.
Subjects & Samples: The generic scaffold is not subject/sample specific but represents an average stomach of subjects used in other studies.
Primary vs derivative data: The primary folder contains the mapping tool provenance data file describing the software environment in place when this dataset was created. The primary folder also contains the settings files which in conjunction with the software information in the provenance file, will reproduce the output files stored in the derivative folder. The derivative folder contains JSON files that are used to generate a webGL visualization of the scaffold on the SPARC portal, as well as the VTK and STL formats of the scaffold.
Code Availability: Scaffold Mapping Tools
Files
1 - 0 of 0 files
About this dataset
Publishing history
Cite this dataset
Tags
References
Is Supplemented by
Lin, M., & Sorby, H. (2024). Publishing Generic Organ Scaffold as a SPARC Dataset v2. https://doi.org/10.17504/protocols.io.6qpvr8q72lmk/v4
