Quantified morphology of the rat vagus nerve
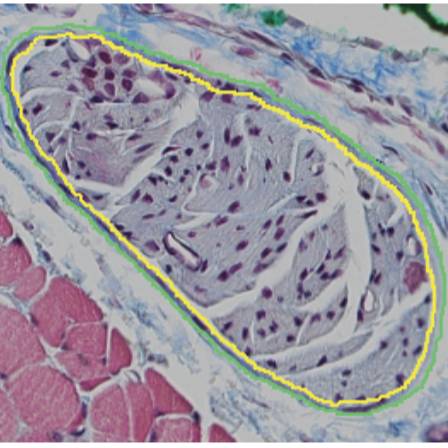
Binary traces from segmentation of cross sections of cervical and subdiaphragmatic rat vagus nerves stained with Masson's trichrome. Quantified effective nerve diameter, effective fascicle diameter, number of fascicles, and perineurium thickness.
Dataset Overview
Study Purpose: To quantify rat vagus nerve morphology
Data Collection: Using micrographs of rat vagus nerve cross sections that were stained with Masson’s trichrome, we quantified effective nerve and fascicle diameters, the number of fascicles, and the perineurium thickness at the cervical and subdiaphragmatic levels. These morphological data provide neural anatomical information, as well as foundational knowledge for computational and preclinical studies of vagus nerve stimulation. The dataset contains TIFF files for 9 left cervical and 9 anterior subdiaphragmatic rat vagus nerve samples, providing segmented traces of inner and outer perineurial boundaries, as well as outer nerve boundaries in multifascicular cases.
Primary Conclusion: None stated
Curator's Notes
Experimental Design: We segmented Masson’s trichrome micrographs of 9 left cervical and 9 anterior subdiaphragmatic rat vagus nerves using Nikon’s NIS-Elements to identify perineurial and neural boundaries. Using the resulting binary traces, we quantified the effective nerve and fascicle diameters, the number of fascicles, and the perineurium thickness.
Completeness: This dataset is part of a larger study "Quantified vagus nerve morphology across species".
Subjects & Samples: Adult male (n=5) and female (n=5) Sprague-Dawley rats were used for this study, 65 - 268 days old.
Primary vs derivative data: Data in the primary and derivative folders are divided by subject number (listed in subject file), then sample number (listed in sample file). The primary folder contains segmentations of rat Masson's trichrome histology (original histological micrographs are in a prior dataset) shown as TIFF images of the binary traces of the fascicles and nerves (loaded and analyzed by Matlab scripts in the "code" folder). The derivative folder contains a .mat file with the morphology metrics output by the code, as well as xml and jp2 versions of the image files; Neurolucida 360 from MBF Bioscience was used to convert the binary traces into xml file format to overlay with the original Masson’s trichrome nd2 micrographs (in prior dataset) and then to convert the images to jp2 extension.
Code Availability: In the "code" folder, Matlab code is used to load the segmented/binary TIFF images, analyze the images, and plot the resulting morphology metrics. Filepaths in the code are set to work as-is, in-place in the dataset.
Files
1 - 0 of 0 files
About this dataset
Publishing history
Cite this dataset
Tags
References
Described by
Pelot, N. A., Goldhagen, G. B., Cariello, J. E., Musselman, E. D., Clissold, K. A., Ezzell, J. A., & Grill, W. M. (2020). Quantified Morphology of the Cervical and Subdiaphragmatic Vagus Nerves of Human, Pig, and Rat. Frontiers in Neuroscience, 14. https://doi.org/10.3389/fnins.2020.601479
Is Supplemented by
A. Pelot, N., B. Goldhagen, G., E. Cariello, J., & M. Grill, W. (2019). SPARC_Duke_PelotGrill_OT2-OD025340_RatVagusNerve_Morphology v1. https://doi.org/10.17504/protocols.io.y6hfzb6
