Population of mock morphological models of vagus nerve stimulation with cuff electrodes for the purpose of studying the effect of fascicle diameter on activation threshold
Computational models and resulting activation thresholds of vagus nerve stimulation for simplified nerve morphologies, built with ASCENT v1.0.3. Modeled electrode designs include monopolar and clinical LivaNova helical cuffs.
Dataset Overview
Study Purpose: To analyze the mechanisms of action of lower activation thresholds for fibers located in smaller fascicles of peripheral nerves through computational modeling .
Data Collection: We used ASCENT v1.0.3 to build 81 3D models of vagus nerves instrumented with cuff electrodes. We simulated activation thresholds for 10 um myelinated fibers across different nerve morphologies, cuff geometries, and stimulation waveforms. We also examined effects of fascicle size and perineurium thickness on distributions of current density and electric potentials.
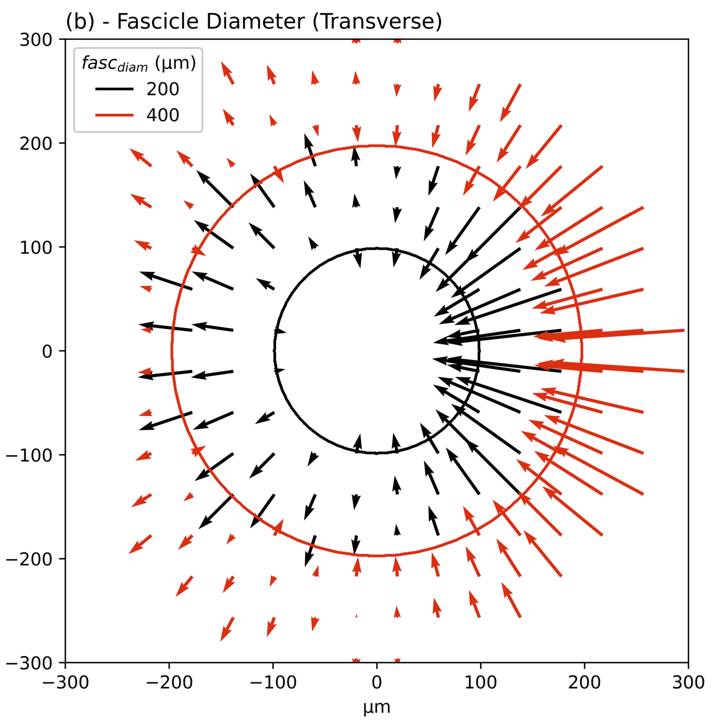
Primary Conclusion: Fibers in larger diameter fascicles have higher activation thresholds than fibers in smaller diameter fascicles due to both thicker perineurium and larger fascicle diameter. The effect of fascicle diameter is larger than the effects of fascicle location or fiber location, but smaller than the effect of fiber diameter. Selective fascicle activation could be achieved with bipolar and tripolar stimulation with small contacts placed on opposite sides of the nerve with a pseudo-monophasic waveform.
Curator's Notes
Experimental Design: Simplified computational models of human vagus nerve stimulation with cuff electrodes.
Completeness: This dataset is complete.
Subjects & Samples: Only virtual subjects were used in this study, which are based on morphological parameters of the human vagus nerve.
Primary vs derivative data: Not applicable. This is a computational study. The files in primary/sub-*/sam-*-sub-*/samples/*/ are ASCENT input configuration files (i.e., sample.json, models/*/model.json) and outputs (i.e., individual fiber activation thresholds models/*/sims/*/n_sims/*/data/outputs/thresh_inner*_fiber*.dat).
Additionally, the files in primary/sub-*/sam-*-sub-*/samples/*/ contain key intermediate files created by ASCENT, including the extracellular potentials applied to the fiber models (models/*/sims/*/n_sims/*/data/inputs/inner*_fiber*.dat) and the stimulation waveform (models/*/sims/*/n_sims/*/data/inputs/waveform.dat).
Code Availability: Run the plotting scripts in the dataset in files/code/ using Python 3.8. The data were generated using ASCENT v1.0.3 (repository; documentation): see README.txt for details.
Files
1 - 0 of 0 files
About this dataset
Publishing history
Cite this dataset
Tags
References
Is Supplement to
Musselman, E. D., Davis, C., Grill, W. M., & Pelot, N. A. (2023). Histology-based computational models of implanted human cervical vagus nerve stimulation with the LivaNova helical cuff electrode [Data set]. In Fibers in Smaller Fascicles Have Lower Activation Thresholds with Cuff Electrodes due to Thinner Perineurium and Smaller Cross-Sectional Area (Version 1). SPARC Portal. https://doi.org/10.26275/KQG4-WBTP
Described by
Davis, C. J., Musselman, E. D., Grill, W. M., & Pelot, N. A. (2023). Fibers in smaller fascicles have lower activation thresholds with cuff electrodes due to thinner perineurium and smaller cross-sectional area. Journal of Neural Engineering, 20(2), 026032. https://doi.org/10.1088/1741-2552/acc42b
