Identification of peripheral neural circuits that regulate heart rate using optogenetic and viral vector strategies part (2)
Data from the innervation of intact mice hearts and the mapping of parasympathetic and sympathetic neural circuits which control heart rate. This data set identifies the cholinergic and noradrenergic neurons which project to the sinoatrial node
Dataset Overview
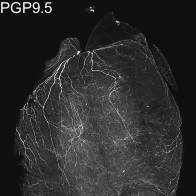
Study Purpose: Here, we developed a clearing-imaging-analysis pipeline to visualize innervation of intact hearts in 3D and employed a multi-technique approach to map parasympathetic and sympathetic neural circuits that control heart rate in mice.
Data Collection: We anatomically and functionally identify cholinergic neurons and noradrenergic neurons in an intrinsic cardiac ganglion and the stellate ganglia, respectively, that project to the sinoatrial node. We also report that the heart rate response to optogenetic versus electrical stimulation of the vagus nerve displays different temporal characteristics and that vagal afferents enhance parasympathetic and reduce sympathetic tone to the heart via central mechanisms
Primary Conclusion: We anatomically and functionally identify cholinergic neurons and noradrenergic neurons in an intrinsic cardiac ganglion and the stellate ganglia, respectively, that project to the sinoatrial node. We also report that the heart rate response to optogenetic versus electrical stimulation of the vagus nerve displays different temporal characteristics and that vagal afferents enhance parasympathetic and reduce sympathetic tone to the heart via central mechanisms.
Curator's Notes
Experimental Design: The mouse was transcardially perfused with phosphate buffered saline (PBS) followed by heparin, prior to the heart tissue being fixed with paraformaldehyde (PFA). The fixed heart was then dehydrated and prepared for iDISCO immunolabeling. Following immunolabeling, the heart was mounted and imaged using confocal or light-sheet microscopy.
Completeness: This data set is part of a larger data set that will also include functional analyses: Identification of peripheral neural circuits that regulate heart rate using optogenetic and viral vector strategies.
Subjects & Samples: This dataset contains data from 12 weeks old (n=35) mice. Samples were derived from the subject's heart.
Primary vs derivative data: Primary data is organized by the subject ID and consists of 1). confocal microscopy to image large tissue volumes and light-sheet microscopy to image entire hearts and when available 2).recordings of in vivo vagus nerve and paravertebral ganglia stimulation. Image data (JPEG2000 and OME-TIFF) in the derivative folder was derived from primary images (.czi). The primary images were converted with 20:1 compression to JPEG2000 (.jp2) with MicroFile+ by MBF Bioscience for web streaming and visualization on the SPARC Data Portal. The primary images were also converted with lossless compression to OME-TIFF (.tif) by MBF Bioscience. Microscopy image metadata is included in the file header of all .jp2 and .tif in the derivative folder, as indicated on the manifest. The derivative folder also contains integrated scaffold files (JSON) for H-4 experimental data that can be viewed and interacted with through the portal viewer.
Important Notes: A web-based interactive visualization of the 3D dataset is available at https://www.ics.uci.edu/~fowlkes/SPARC/volume/. A detailed description of procedures and findings is published in Nature Communications doi: 10.1038/s41467-019-09770-1
Files
1 - 0 of 0 files
About this dataset
Publishing history
Cite this dataset
Tags
References
Is Supplemented by
Rajendran, P. (2019). iDISCO clearing of mouse heart v1. https://doi.org/10.17504/protocols.io.x3sfqne
Described by
Rajendran, P. S., Challis, R. C., Fowlkes, C. C., Hanna, P., Tompkins, J. D., Jordan, M. C., Hiyari, S., Gabris-Weber, B. A., Greenbaum, A., Chan, K. Y., Deverman, B. E., Münzberg, H., Ardell, J. L., Salama, G., Gradinaru, V., & Shivkumar, K. (2019). Identification of peripheral neural circuits that regulate heart rate using optogenetic and viral vector strategies. Nature Communications, 10(1). https://doi.org/10.1038/s41467-019-09770-1
