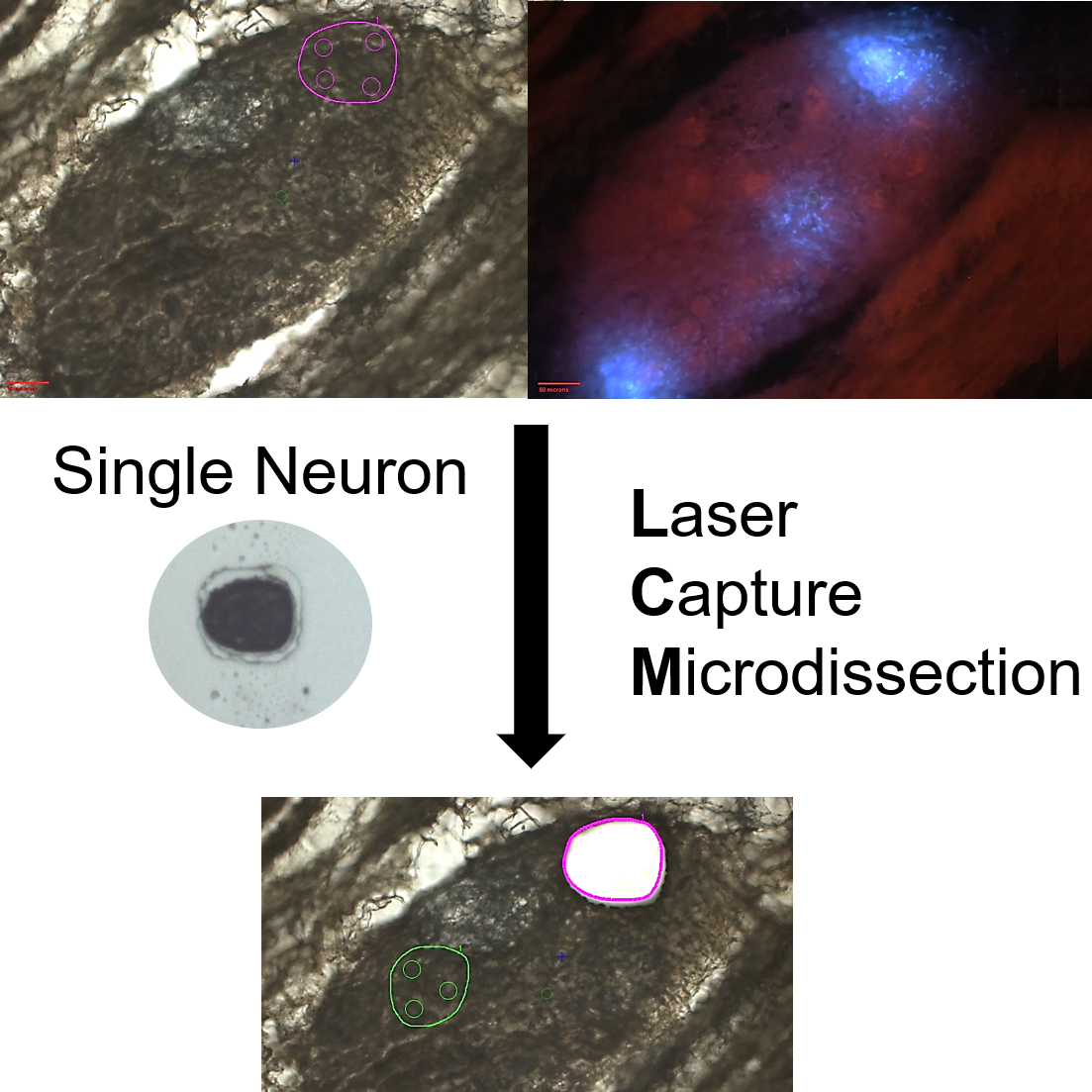
Acquisition of single neurons and regional neuronal samples from the porcine right atrial ganglionic plexus (RAGP) through laser capture microdissection
Transcriptional diversity of single neurons in the porcine RAGP. High throughput qPCR data for over 400 single neurons assayed for hundreds of genes across 4 pig RAGP.
Dataset Overview
Study Purpose: The purpose of this study was to create a comprehensive atlas of the cardiac ICN in porcine Right atrial Ganglionic Plexus (RAGP) at a cellular level providing gene expression profiles of cardiac neurons at single-cell resolution. We developed an approach to appreciate the 3D organization of the intracardiac neurons, ICN, while at the same time permitting single-cell transcriptomics and connectomics.
Data Collection: This dataset contains images that track the collection of single neurons and regional neuronal samples through Arcturus Laser Capture Microdissection (LCM), totaling 1215 total samples collected across 4 RAGP (415 single neurons went on to be used for HT-qPCR, 142 regional neuronal samples went on to RNAseq).
Primary Conclusion: Through serial cryostat sectioning of a cryopreserved heart with imaging of serially collected and stained sections, it is possible to collect single neuron and regional neuronal samples through laser capture microdissection. The transcriptional profiles of these samples can be determined down to single-cell resolution and mapped back into the 3D context generated by stacking the serial images and using the images taken to track their precise locations.
Curator's Notes
Experimental Design: The porcine heart was injected with FastBlue, a retrograde neuronal tracer, into the SAN before sacrifice, allowing the distinction between SAN-connecting neurons and non SAN-connecting neurons based on whether they stain for just cresyl violet (non SAN-connecting), or cresyl violet and FastBlue (SAN-connecting). Overview images were taken of the complete organ as well as at several different levels of magnification both before the samples were lifted and after. Such sample tracking later allowed us to map the transcriptomics of the assayed samples back to their 3D locations.
Completeness: This dataset is a part of a larger study: Spatially Tracked Single Neuron Anatomical and Molecular Profiling of the Pig Right Atrial Ganglionic Plexus.
Subjects & Samples: Male (n=2) and female (n=2) adolescent Yucatan pigs were used for this study to collect Arcturus Laser Capture Microdissection (LCM), totaling 1215 of single-neuron samples.
Primary vs derivative data: Each subfolder within the "primary" folder contains three items: Folders of sample acquisition images, a xlsx file detailing the sample collection information, and a xlsx file explaining the labeling convention for each sample. Image data (JPEG2000 and OME-TIFF) in the derivative folder was derived from primary images (.jpeg). The primary images were converted with 20:1 compression to JPEG2000 (.jp2) by MBF Bioscience for web streaming and visualization on the SPARC Data Portal. The selection of primary images were also converted with lossless compression to OME-TIFF (.tif) by MBF Bioscience. Microscopy image metadata is included in the file header of all .jp2 and .tif in the derivative folder, as indicated on the manifest.
Important Notes: This dataset is currently undergoing image registration and will be updated once this process is complete.
Files
0 - 0 of 0 files
About this dataset
Publishing history
Cite this dataset
Tags
References
Is Supplemented by
Robbins, S., Achanta, S., & Vadigepalli, R. (2021). Laser Capture Microdissection (LCM) and 3D Sample Tracking Protocol. https://doi.org/10.21203/rs.3.pex-927/v1
Robbins, S., Vadigepalli, R., & Schwaber, J. (2021). Cryosectioning, block face imaging and Nissl staining fluorescently labeled pig heart. https://doi.org/10.21203/rs.3.pex-928/v1
