Computational analysis of the human sinus node action potential - Model development and effects of mutations
CellML version of the Fabbri et al. 2017 mathematical model of the spontaneous electrical activity of a human sinoatrial node (SAN) pacemaker cell
Dataset Overview
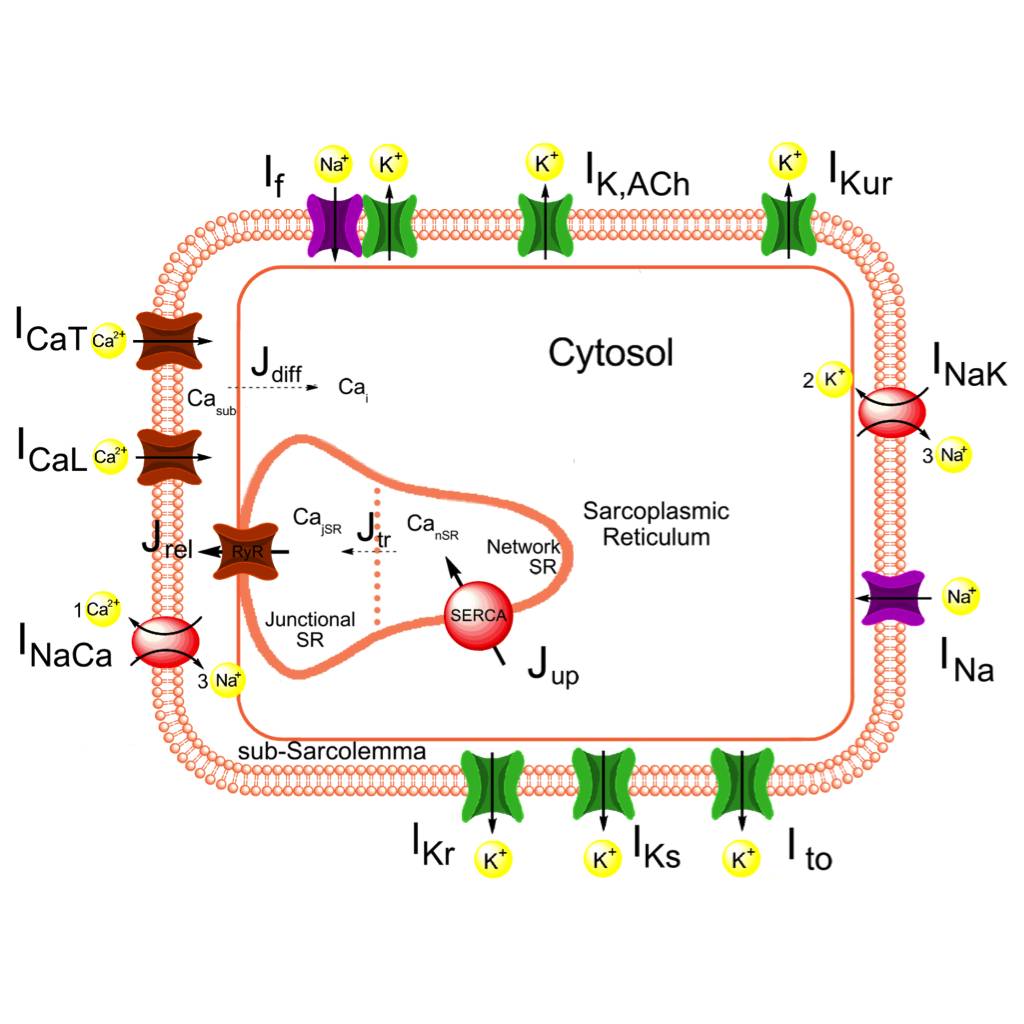
Study Purpose: Fabbri et al. constructed a comprehensive mathematical model of the spontaneous electrical activity of a human sinoatrial node (SAN) pacemaker cell, starting from the recent Severi–DiFrancesco model of rabbit SAN cells.
Data Collection: The model is based on electrophysiological data from isolated human SAN pacemaker cells and closely matches the action potentials and calcium transient that were recorded experimentally.
Primary Conclusion: Simulated ion channelopathies explain the clinically observed changes in heart rate in corresponding mutation carriers, providing independent qualitative validation of the model. The model shows that the modulatory role of the ‘funny current’ (If) in the pacing rate of human SAN pacemaker cells is highly similar to that of rabbit SAN cells, despite its considerably lower amplitude. The model may prove useful in the design of experiments and the development of heart‐rate modulating drugs.
Curator's Notes
Experimental Design: This is a computational study.
Completeness: Complete.
Subjects & Samples: Not applicable. This is a computational study.
Primary vs derivative data: Not applicable. This is a computational study, and no primary data was collected. Links to instrumented versions of the CellML models and metadata are provided in the dataset_description file.
Code Availability: CellML version of the Fabbri et al. 2017 model.
Files
1 - 0 of 0 files
About this dataset
Publishing history
Cite this dataset
Tags
References
Described by
Fabbri, A., Fantini, M., Wilders, R., & Severi, S. (2017). Computational analysis of the human sinus node action potential: model development and effects of mutations. The Journal of Physiology, 595(7), 2365–2396. Portico. https://doi.org/10.1113/jp273259
