Micro Computed Tomography (Micro-CT) imaging of iodine-stained rat stomachs from full to empty
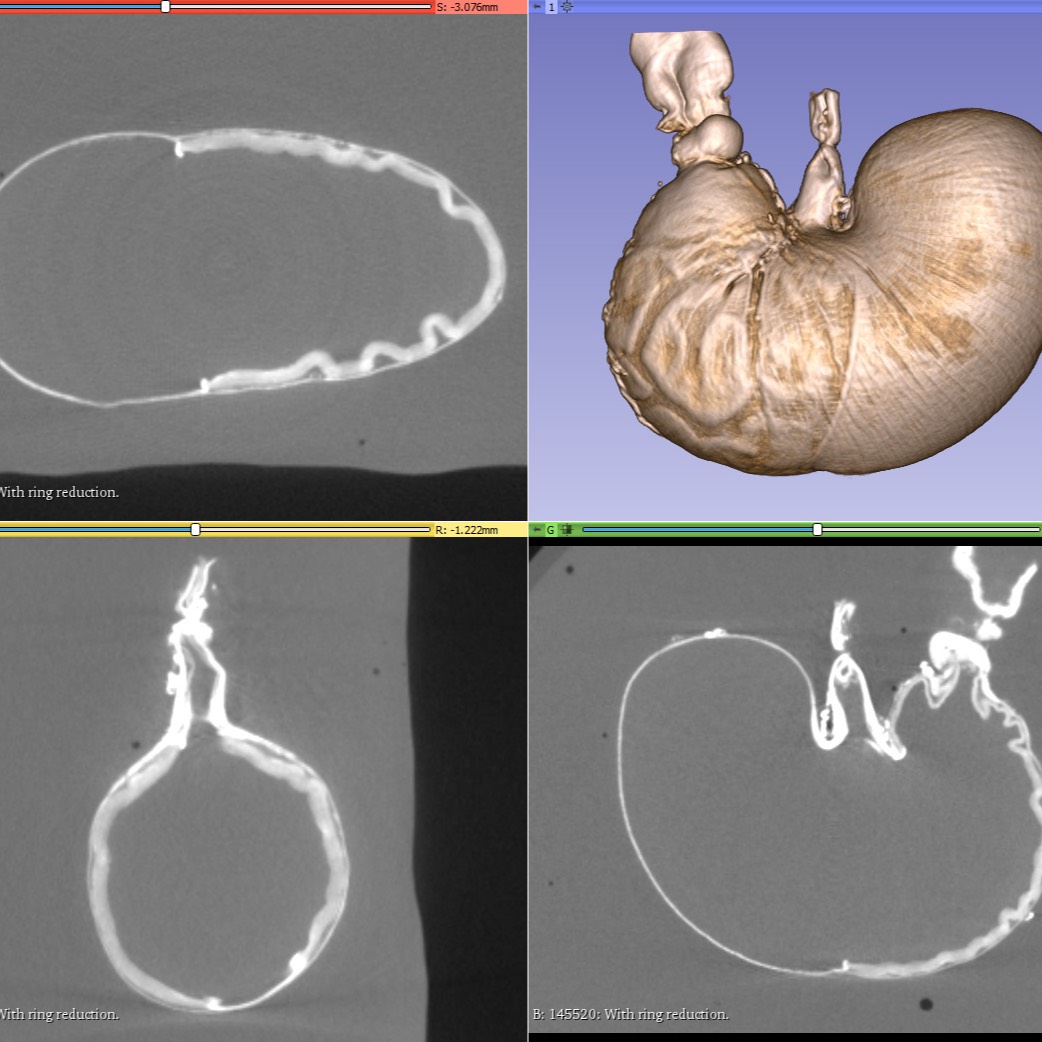
Micro Computed Tomography (Micro-CT) images of the stomachs stained with an aqueous solution of I2/KI. Micro-CT images were saved in DICOM format. Stomachs were scanned both before and after flushing of stomach contents.
Dataset Overview
Study Purpose: As part of the SPARC research, we are interested in understanding how the rat stomach's dimensions vary as a function of feeding and elapsed time since feeding. Our measurements enable generating a 4D model of the rat stomach integrating anatomical details with 2D maps of neurite distributions and other anatomical distributions. This model can then be compared with functional behavior mapping results or used to predict preferred stimulation locations for gastric electrical stimulation experiments.
Data Collection: To provide contrast in micro-CT, the stomachs were stained with an aqueous solution of I2/KI. Micro-CT images were saved in DICOM format. Stomachs were scanned both before and after flushing of stomach contents. 3D Slicer was used to quantify both total stomach volume and the volumes of subcompartments of the stomach. A subset was segmented using Neurolucida 360 software to provide the elements needed to create a stomach scaffold, and those XML files are included.
Primary Conclusion: None stated
Curator's Notes
Experimental Design: Whole rat stomachs were dissected and prepared for micro-CT imaging to enable a 3D organ description. Iodine was used to stain the tissue post-perfusion. By varying the meal size and the delay time between feeding and perfusion, researchers determined how stomach volume changes, both overall and for stomach compartments. The DICOM file output was used to generate a 3D scaffold of the stomach for modeling purposes.
Completeness: this dataset is complete
Subjects & Samples: Male Sprague Dawley rats (n = 46) weighing 228‐330 g were used in the study.
Primary vs derivative data: The data set includes DICOM and NRRD ("nearly raw raster data") files (in tar.bz2 directories, 4.1 GB) in bz2 compressed tape archive file for each sample organized in primary data folder by subject's name. 34 XML files (Neurolucida segmentation - about 200 MB total) and an Excel (.XLSX) spreadsheet detailing volume measurements from 3D Slicer segmentation for a subset of subjects can be found in the derivative folder.
Important Notes: A detailed 3D Slicer Segmentation Protocol used in this study is included as a World document in the docs folder.
Files
1 - 0 of 0 files
About this dataset
Publishing history
Cite this dataset
Tags
References
Is Supplemented by
Jaffey, D., Powley, T., & Chesney, L. (2019). Micro-CT imaging of iodine-stained rat stomach v1. https://doi.org/10.17504/protocols.io.95ih84e
